Using complete.cases() with my actual data is too aggressive and removes too much data. This seems to be due to the NA's, which are only in some SOA conditions. The error is: Error in summarise_impl(.data, dots) :Ĭolumn `Peak_Latency` must be length 1 (a summary value), not 0 I am running into a problem because some subjects have missing data (NA's). Because R makes dealing with this missing data so easy is another reason it is so often used in statistical analysis.I am working with a data set of changes over time and need to calculate the time at which the peak change occurs. That makes all the difference.ĭealing with missing data from a data set is critical to proper data science. Selection helpers can be used in functions like dplyr::select() or. length() doesnt take na.rm as an option, so one way to work. startswith(match, ignore.case TRUE, vars NULL) endswith(match, ignore.case. To have R ignore these, we tell the mean function to remove the NA s. 
The second and third examples are identical except that in the first one na.rm = FALSE, and na.rm = TRUE in the other. If there are NAs in the data, you need to pass the flag na.rmTRUE to each of the functions. summarize(groupby(demdf, riagendr), meanage mean(ridageyr, na.rm TRUE)). Consider the R code and its output below: datagroupNA <- data, lapply (.SD, mean), Summarize data.table by group by group datagroupNA Print summarized data.table. This example demonstrates what happens when we do not actively avoid NA values when summarizing a data.table in R. You can install it from CRAN with: install.packages ('dplyr') You can see a full list of changes in the release notes. Example 1: Summarize data.table without Removing NA. Note the NA in row 3 column b, this shall be the missing data set for these examples. dplyr Romain Francois We’re happy to announce the release of dplyr 1.0.4, featuring: two new functions ifall () and ifany (), and improved performance improvements of across (). To start our examples, we need to set up a dataframe to work from. However, when na.rm is FALSE, then it returns NA from the calculation being done on the entire row or column. but it ignores the 'of all columns' in this question. You can use multiple mean statements in dplyr::summarize like this. When na.rm is TRUE, the function skips over any NA values. In your original answer and in 'Edit2' how would you enter the na.rm TRUE argument into the mean function. They include colSums(), rowSums(), colMeans() and rowMeans(). You can simply use mean from hablar that has na.rm T as default: library (hablar) df > summariseall (mean) var1 var2 var3 var4 1 6.666667 4.666667 1 4. It is simply a parameter used by several dataframe functions. 
It is neither a function nor an operation. with vars(): starwars > summariseat(vars(height:mass), mean, na.rm TRUE). When using a dataframe function na.rm in r refers to the logical parameter that tells the function whether or not to remove NA values from the calculation. Grouping variables covered by implicit selections are silently ignored by. The two remove NA values in r is by the na.omit() function that deletes the entire row, and the na.rm logical perimeter which tells the function to skip that value. While this may be okay sometimes in other cases you need a number. R: Combine columns ignoring NAs - Stack Overflow WebThis example demonstrates what happens when we do not actively avoid NA values when summarizing a. If you include the NA value in a calculation it will result in an NA value. 
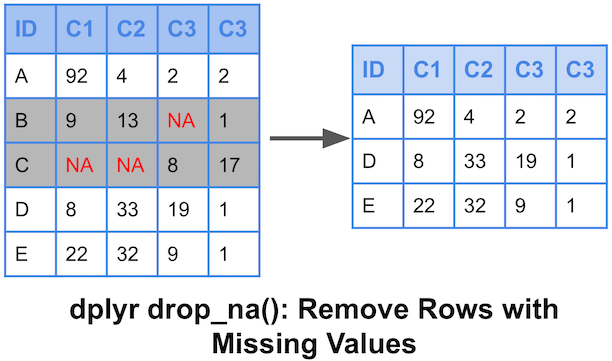
So, somehow it needs to be removed from the calculations to get a meaningful value. one way of dealing with missing data with dataframe functions is through the na.rm logical perimeter.īecause the NA value is a placeholder and not an actual numeric value, it cannot be included in calculations. Method 1: Remove Rows with NA Values in Any Column library(dplyr) remove rows with NA value in any column df > na.omit() Method 2: Remove Rows with NA Values in Certain Columns library(dplyr) remove rows with NA value in 'col1' or 'col2' df > filterat (vars (col1, col2), allvars (is.na(. While the cbind() function will accept data containing NA, it does produce a warning. It is accepted by ame() without difficulty. Another the na.omit() function deletes any rows in the dataframe containing missing data in R missing data is designated by NA so that it can be detected easily. One way is the is.na() function involves simply detecting it. There are several ways to deal with missing data in r. Sometimes, things beyond your control can cause gaps in the data. In a lab, you can control the quality of the data, but the real world does not work so nicely. In base R, use na.omit() to remove all observations with missing data on ANY variable in the dataset, or use subset() to filter out cases that are missing. If you have ever done any research involving real-world measurements, then you know that the data is not always neat and tidy. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
